sctrial — Participant-Level Differential Analysis for Longitudinal Single-Cell Experiments#


sctrial is a Python package for participant-level differential analysis of longitudinal single-cell RNA-seq data. Built on AnnData, it aggregates cell-level measurements to participant-level replicates and applies design-specific statistical methods — difference-in-differences, paired contrasts, and cross-sectional comparisons — so that inference reflects the actual number of independent biological units in the study.
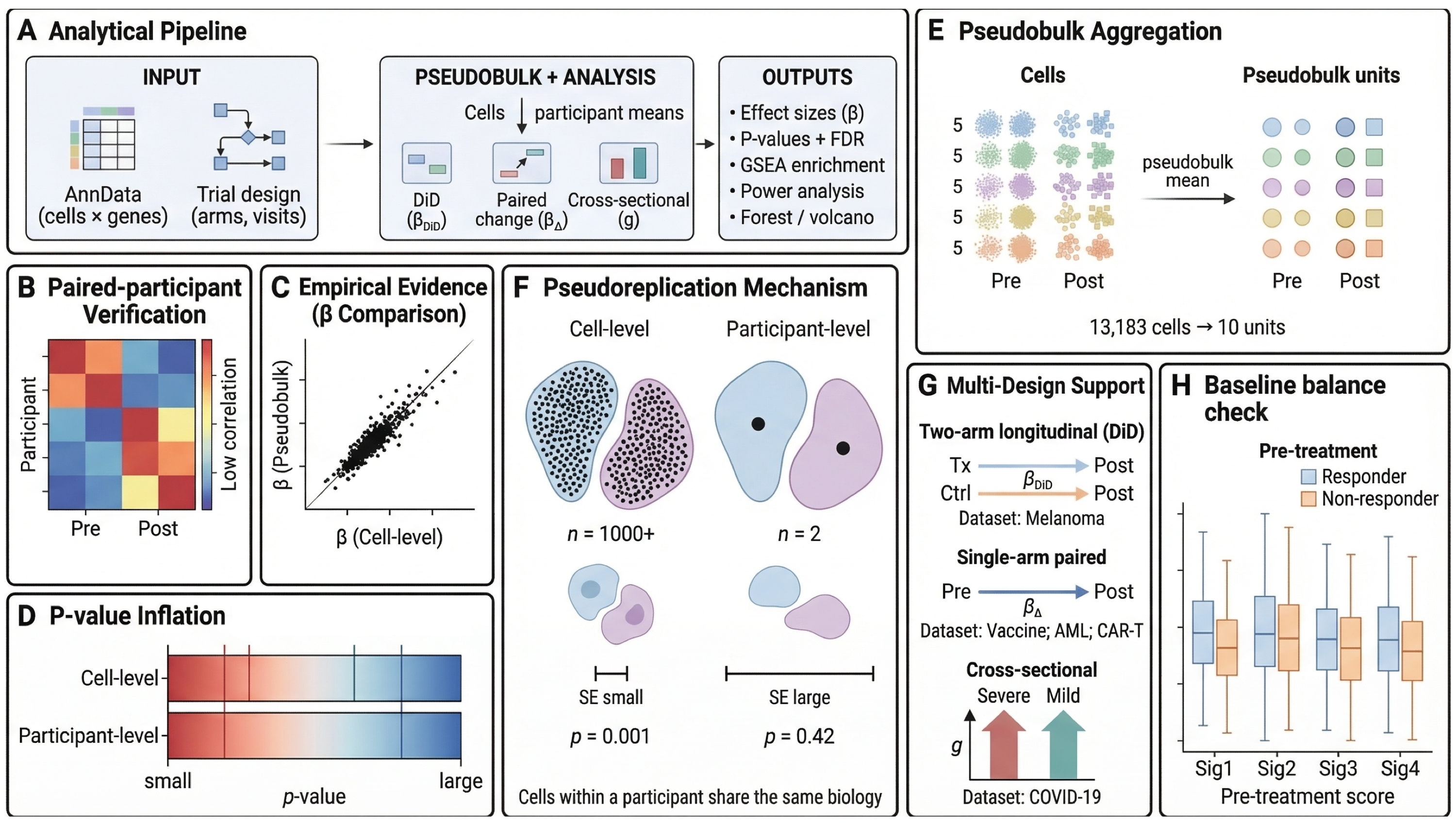
Key Applications#
Difference-in-Differences (DiD) — Participant fixed-effects regression with wild cluster bootstrap inference for treatment effect estimation across arms and timepoints.
Paired Contrasts — Within-arm pre→post comparisons with paired statistical tests (Wilcoxon, t-test) and effect sizes (Cohen’s d, Hedges’ g, log₂ fold-change).
Between-Arm Tests — Cross-sectional comparisons between treatment and control arms at fixed timepoints with proper pseudobulk replication.
Abundance Analysis — Cell-type composition changes across conditions using proportion-based statistics and participant-level aggregation.
Gene Set Enrichment Analysis — GSEA on DiD-ranked gene lists with support for Hallmark, KEGG, Reactome, and custom gene set collections.
Power Analysis — Sample size calculations, power curves, and minimum detectable effect sizes for planning single-cell clinical studies.
Getting Started#
Install sctrial and run your first analysis:
pip install sctrial
See the Installation guide for optional extras (plotting, GSEA, Bayesian modules), then follow the Quickstart for a complete walkthrough. For end-to-end analyses on real clinical datasets, explore the Tutorials.
Citing sctrial#
If you use sctrial in your research, please cite:
Vasanthakumari P, Valencia I, Aghmiouni MR, Magana B, Omar M. sctrial: Participant-Level Differential Analysis for Longitudinal Single-Cell Experiments. bioRxiv 2026. doi:10.64898/2026.04.02.716219.
@article{vasanthakumari2026sctrial,
title = {sctrial: Participant-Level Differential Analysis for Longitudinal Single-Cell Experiments},
author = {Vasanthakumari, Priyanka and Valencia, Itzel and Aghmiouni, Maryam R. and Magana, Bryan and Omar, Mohamed},
journal = {bioRxiv},
year = {2026},
doi = {10.64898/2026.04.02.716219},
url = {https://www.biorxiv.org/content/10.64898/2026.04.02.716219v1}
}