New Preprint: sctrial — Participant-Level Differential Analysis for Longitudinal Single-Cell Experiments
April 6, 2026
We are excited to share our new preprint, “sctrial: Participant-Level Differential Analysis for Longitudinal Single-Cell Experiments,” now available on bioRxiv.
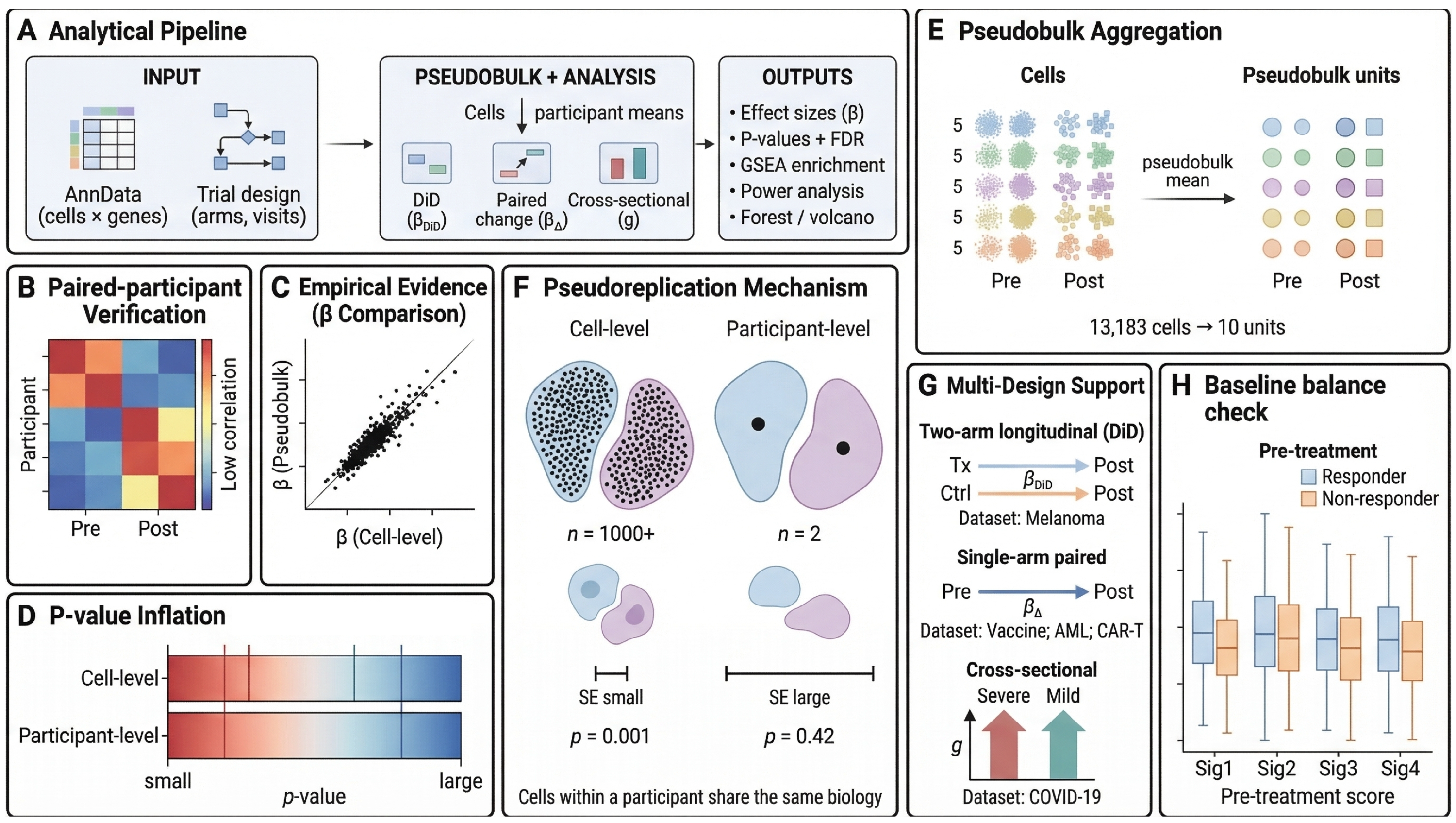
Longitudinal single-cell RNA sequencing studies in clinical trials and translational cohorts offer a powerful view of treatment response, disease progression, and cellular dynamics. Their hierarchical structure, however, poses a major inferential challenge: thousands of cells are measured within the same participants across repeated time points, but the participant — not the cell — is the true unit of biological replication. Conventional cell-level workflows can therefore yield inflated significance and misleading confidence in reported associations.
sctrial is an open-source Python package for repeated-measures single-cell studies that uses design-specific participant-level estimands, including difference-in-differences for two-group longitudinal comparisons, and small-cluster-aware uncertainty quantification. In simulation benchmarks using a hierarchical gamma-Poisson generative model, sctrial maintained well-calibrated error rates in mixed-signal gene panels where established multi-subject methods showed inflated false positive rates among unaffected genes.
We applied sctrial to five independent datasets spanning melanoma immunotherapy, COVID-19 severity, BNT162b2 vaccination, AML chemotherapy, and CAR-T therapy. Across these studies, sctrial identified immune programs whose direction and magnitude differed across therapeutic and disease contexts, while benchmarking analyses showed that many associations highlighted by conventional cell-level workflows were attenuated or no longer supported when inference was performed at the participant level.
The package is available on PyPI, conda-forge, and GitHub, with full documentation at omar-lab.com/sctrial.